|
Size: 4554
Comment:
|
Size: 4379
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 27: | Line 27: |
| [[attachment:example_scans.tif|Examples of different quality scans (click to download TIFF image):|&do_get]] | {{attachment:example_scan_A.jpg}} {{attachment:example_scan_B.jpg}} {{attachment:example_scan_C.jpg}} |
| Line 41: | Line 43: |
| 1. Read the introductory material on FreeSurfer from past lectures: * [[attachment:FSL_anatomical_stream.pdf|slides 1 (BF)]] * [[attachment:fsintro.pdf|slides 2 (DG)]] * [[attachment:fsrecon.pdf|slides 3 (DG)]] |
1. Read the links to slides marked as type 'talk' in the tutorials: [[FsTutorial|click here]] |
FreeSurfer Beginners Guide
FreeSurfer is a freely available software package developed by investigators at the Athinoula A. Martinos Center for Biomedical Imaging used for a number of procedures including:
- Creation of computerized models of the brain from magnetic resonance imaging (MRI) data.
- Processing of functional magnetic resonance imaging (fMRI) data.
- Measuring a number of morphometric properties of the brain including cortical thickness and regional volumes.
- Intersubject averaging of structural and functional data using a procedure that aligns individuals based on their cortical folding patterns for optimal alignment of homologous neural regions.
Machine Requirements
To run FreeSurfer, you will need either a PC running Linux or a Macintosh running OS X.
FreeSurfer consumes a lot of processor time, memory resources and disk space, so it is recommended to run FreeSurfer on as powerful a machine as you have available. For example, at MGH we typically run Linux CentOS 5 on 2.5GHz quad processor Intel Xeon with 4 to 8 GB of DDR SDRAM, and 500GB of disk space.
See SystemRequirements for more info.
Data Requirements
The processing procedures for the creation of cortical models require good quality T1 weighted MRI data, such as a Siemens MPRAGE (examples of appropriate Siemens scanner protocols) or GE SPGR sequence with approximately 1mm3 resolution (although a variety of quality data sets can be processed with additional manual intervention). Thickness should not exceed 1.5mm (~1mm^3 is ideal). The best FreeSurfer processing results come from scans having excellent gray/white matter contrast.
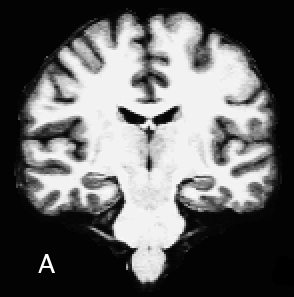
A - excellent GM/WM contrast, acquired with MP-RAGE pulse-sequence protocol
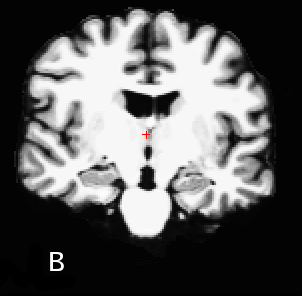
B - very good GM/WM contrast, acquired with MP-RAGE pulse-sequence protocol
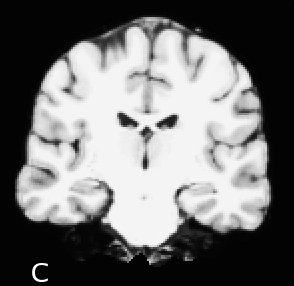
C - good GM/WM contrast, acquired with SPGR pulse-sequence protocol; not as good GM/WM contrast as the MP-RAGE acquisitions; low bandwidth causes some temporal lobe artifacts (brightening of the gray matter)
These example scans, captured from FreeSurfer's tkmedit display, are slices from the brainmask.mgz file, created by the -autorecon1 stage of FreeSurfer (requiring about 20 minutes of processing time). This means that some amount of intensity normalization has occurred, but the contrast differences between image examples are easier to see, compared to the original (orig.mgz) volume. Refer to the tutorial sections 'Using Control Points to Fix Intensity Normalization' and 'Fixing Common Geometric Inaccuracies in White Matter Surfaces' for examples of problem areas resulting from scan deficiencies.
Getting Started
There is a variety of documentation about the use of FreeSurfer contained in the FreeSurfer wiki including installation of the software, tutorials, sample data, and work flows providing step by step guides to performing specific tasks. To get started, we suggest you:
Read the links to slides marked as type 'talk' in the tutorials: click here
Read the background material: click here
Install FreeSurfer: click here
Download the sample data set: click here
Follow the cortical reconstruction tutorial to create cortical models: click here
Follow the reconstruction workflow page: click here
Peruse the wiki to get a fuller knowledge of all of the available processing procedures in the FreeSurfer software package
An active e-mail list is available to answer specific questions about processing procedures.
There is also a FAQ
