|
Size: 1052
Comment:
|
Size: 2008
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 1: | Line 1: |
| [[fswiki|AnatomiCuts]] | |
| Line 4: | Line 3: |
| This method finds corresponding clusters across subjects. Currently, only one-to-one cluster's correspondences are available using the Hungarian algorithm. | This method finds corresponding clusters across subjects. Currently, only one-to-one cluster's correspondences are available using the Hungarian algorithm. Importantly, by using our anatomical similarity metric, correspondences are found without the need of registration comparing clusters of streamlines from each subject's native space. {{attachment:hungarian_babies.png||height="360px"}}{ |
| Line 15: | Line 16: |
| Where |
|
| Line 16: | Line 19: |
| Line 17: | Line 21: |
| Line 18: | Line 23: |
| Line 19: | Line 25: |
| Line 20: | Line 27: |
| Line 21: | Line 29: |
| Line 23: | Line 32: |
== Output == The output will be a csv file: {{{ Subject A, Subject B 100000,11111 1010101, 100010 }}} Where cluster 100000.trk from Subject A corresponds to cluster 11111.trk from Subject B |
|
| Line 24: | Line 49: |
V. Siless, J. Y. Davidow, J. Nielsen, Q. Fan, T. Hedden, M. Hollinshead, C. V. Bustamante, M. K. Drews, K. R. A. Van Dijk, M.A. Sheridan, R. L. Buckner, B. Fischl, L. Somerville, and A. Yendiki. 2017. “Registration-free analysis of diffusion MRI tractography data across subjects through the human lifespan.” V. Siless, K. Chang, B. Fischl, and A. Yendiki. 2018. “AnatomiCuts: Hierarchical clustering of tractography streamlines based on anatomical similarity.” NeuroImage, 166, Pp. 32-45. |
AnatomiCuts correspondences
This method finds corresponding clusters across subjects. Currently, only one-to-one cluster's correspondences are available using the Hungarian algorithm. Importantly, by using our anatomical similarity metric, correspondences are found without the need of registration comparing clusters of streamlines from each subject's native space.
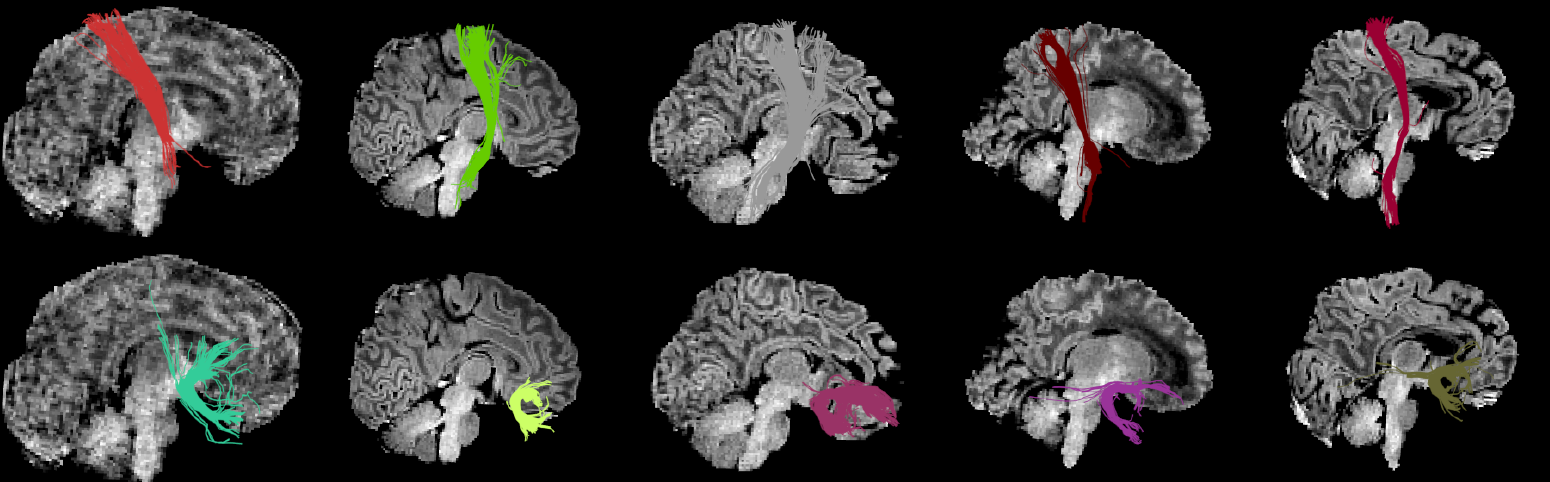 {
{
The Hungarian algorithm
The Hungarian algorithm finds corresponding clusters between two subjects.
AnatomiCuts_correspondences -s1 segmentation1.nii.gz -s2 segmentation2.nii.gz -c numClusters -h1 clusteringPath1 -h2 clusteringPath2 -m metric -o output.csv
Where
-s1 the segmentation to be used for anatomical similarity in subject one.
-s2 the segmentation to be used for anatomical similarity in subject one.
-h1 the path to the AnatomiCuts folder to be used for subject1.
-h2 the path to the AnatomiCuts folder to be used for subject2.
-m metric to be used: labels (anatomical similarity) or euclid (euclidean similarity).
-sym (under development) this flag will mirror the segmentation in subject 2 to find between hemisphere correspondences.
-o output csv file
Output
The output will be a csv file:
Subject A, Subject B 100000,11111 1010101, 100010
Where cluster 100000.trk from Subject A corresponds to cluster 11111.trk from Subject B
References
V. Siless, J. Y. Davidow, J. Nielsen, Q. Fan, T. Hedden, M. Hollinshead, C. V. Bustamante, M. K. Drews, K. R. A. Van Dijk, M.A. Sheridan, R. L. Buckner, B. Fischl, L. Somerville, and A. Yendiki. 2017. “Registration-free analysis of diffusion MRI tractography data across subjects through the human lifespan.”
V. Siless, K. Chang, B. Fischl, and A. Yendiki. 2018. “AnatomiCuts: Hierarchical clustering of tractography streamlines based on anatomical similarity.” NeuroImage, 166, Pp. 32-45.
