| Deletions are marked like this. | Additions are marked like this. |
| Line 1: | Line 1: |
| This process uses the command ["mri_fill"] to create a binarized volume. The input is the mri/wm volume and the output is the mri/filled volume. First, the cutting planes are determined to separate the hemispheres from each other and the pons. Then a seed point is located and used to fill each hemisphere. The rh will be filled with a value of 255, and the lh with a value of 127. | This process uses the command [[mri_fill]] to create a binarized volume. The input is the mri/wm volume and the output is the mri/filled volume. First, the cutting planes are determined to separate the hemispheres from each other and the pons. Then a seed point is located and used to fill each hemisphere. The rh will be filled with a value of 255, and the lh with a value of 127. |
| Line 5: | Line 5: |
| 1. Open the subject's (subj) wm volume up in ["tkmedit"]. | 1. Open the subject's (subj) wm volume up in [[tkmedit]]. |
| Line 7: | Line 7: |
| ["tkmedit"] subj wm | [[tkmedit]] subj wm |
| Line 15: | Line 15: |
| attachment:CCcor.jpg attachment:CChor.jpg attachment:CCsag.jpg | {{attachment:CCcor.jpg}} {{attachment:CChor.jpg}} {{attachment:CCsag.jpg}} |
| Line 19: | Line 19: |
| ["recon-all"] -cc-xyz .... | [[recon-all]] -cc-xyz .... |
| Line 23: | Line 23: |
| ["mri_fill"] -C .... | [[mri_fill]] -C .... |
| Line 25: | Line 25: |
| Alternatively seed point can be given in voxel coordinates under the command line ["mri_fill"] : | Alternatively seed point can be given in voxel coordinates under the command line [[mri_fill]] : |
| Line 35: | Line 35: |
| attachment:Pcor.jpg attachment:Phor.jpg attachment:Psag.jpg | {{attachment:Pcor.jpg}} {{attachment:Phor.jpg}} {{attachment:Psag.jpg}} |
| Line 39: | Line 39: |
| attachment:RHcor.jpg attachment:RHhor.jpg attachment:RHsag.jpg | {{attachment:RHcor.jpg}} {{attachment:RHhor.jpg}} {{attachment:RHsag.jpg}} |
| Line 43: | Line 43: |
| attachment:LHcor.jpg attachment:LHhor.jpg attachment:LHsag.jpg | {{attachment:LHcor.jpg}} {{attachment:LHhor.jpg}} {{attachment:LHsag.jpg}} |
| Line 46: | Line 46: |
|
[[BR]] [[BR]] ["mri_fill"] -C MNIcoords -P MNIcoords -rh MNIcoords -lh MNIcoords $SUBJECTS_DIR/subj/mri/wm $SUBJECTS_DIR/subj/mri/filled;inflate_subject-rh subj;inflate_subject-lh subj [[BR]] [[BR]] |
<<BR>> <<BR>> [[mri_fill]] -C MNIcoords -P MNIcoords -rh MNIcoords -lh MNIcoords $SUBJECTS_DIR/subj/mri/wm $SUBJECTS_DIR/subj/mri/filled;inflate_subject-rh subj;inflate_subject-lh subj <<BR>> <<BR>> |
| Line 51: | Line 51: |
|
[[BR]][[BR]] ["recon-all"] -stage2 -cc-xyz MNIcoords -pons-xyz MNIcoords -rh-xyz MNIcoords -lh-xyz MNIcoords -subjid subj [[BR]][[BR]] |
<<BR>><<BR>> [[recon-all]] -stage2 -cc-xyz MNIcoords -pons-xyz MNIcoords -rh-xyz MNIcoords -lh-xyz MNIcoords -subjid subj <<BR>><<BR>> |
This process uses the command mri_fill to create a binarized volume. The input is the mri/wm volume and the output is the mri/filled volume. First, the cutting planes are determined to separate the hemispheres from each other and the pons. Then a seed point is located and used to fill each hemisphere. The rh will be filled with a value of 255, and the lh with a value of 127.
Occasionally, manual intervention is required to define the cutting planes and seed points. To do so, follow these steps:
Open the subject's (subj) wm volume up in tkmedit.
tkmedit subj wm
View the MniTalairach coordinates: View->Information->MNI Coordinates
- Select the Corpus Callosum point, on the mid-posterior portion of it. Example: Corpus Callosum
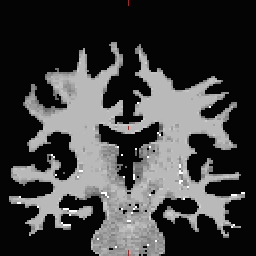
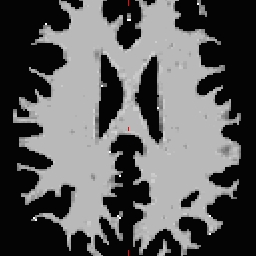
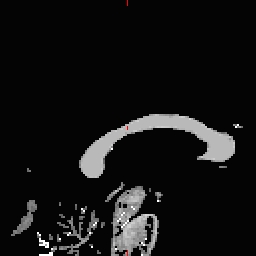
- Copy the MNI coordinates to the command line for either
recon-all -cc-xyz .... or in
mri_fill -C ....
Alternatively seed point can be given in voxel coordinates under the command line mri_fill :
-CV <x> <y> <z> - the voxel coords of the seed for the corpus callosum
-PV <x> <y> <z> - the voxel coords of the seed for the pons
- Do this for all of the other cutting plane points and seed points. Examples follow: Pons
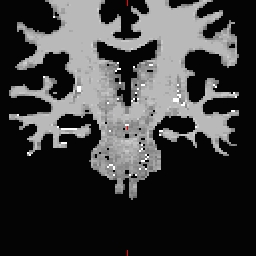
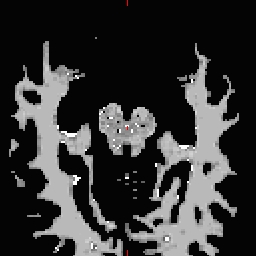
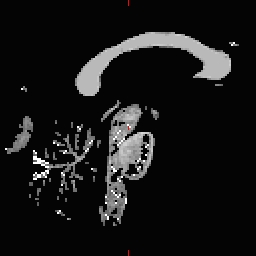 RH seed
RH seed 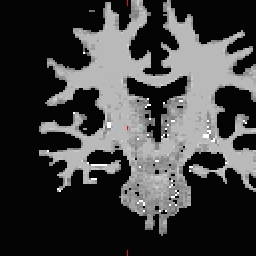
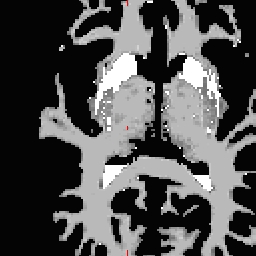
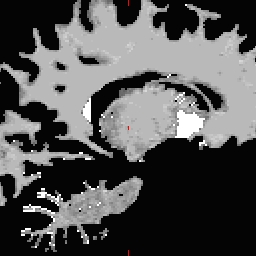 LH seed
LH seed 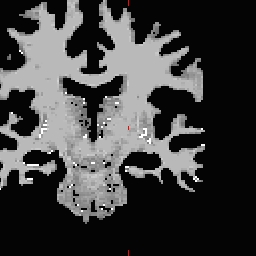
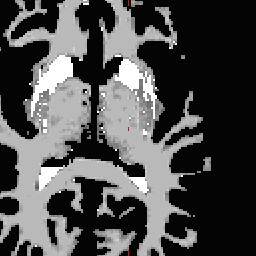
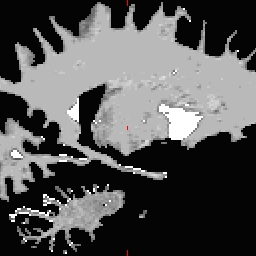
- Not all points need to be defined, any range from none to all four may be defined. To inflate:
mri_fill -C MNIcoords -P MNIcoords -rh MNIcoords -lh MNIcoords $SUBJECTS_DIR/subj/mri/wm $SUBJECTS_DIR/subj/mri/filled;inflate_subject-rh subj;inflate_subject-lh subj
or, the equivalent in recon-all:
recon-all -stage2 -cc-xyz MNIcoords -pons-xyz MNIcoords -rh-xyz MNIcoords -lh-xyz MNIcoords -subjid subj
NOTE: These command line(s) need to be run EVERYTIME the subject's surfaces are reinflated
